Postprocessing MCMC draws
Each MCMC run generates posterior draws (samples) in .ext and .iph, as described in output files. In this article we show how to postprocess these draws in R.
Consider the following example. The model uses ITS to position the initial draws, followed by NUTS sampler that draws.
|
|
The raw output file contains the draws for THETA, OMEGA, and SIGMA:
|
|
Note that the delim=q option for $EST METHOD=NUTS effectively
saves .ext draws as a csv file and adds quotation to the column names.
The NITER=1000 option requests 1000 draws. Their summary and
diagnostic information is printed on lines with dummy iteration id's "-100000000x".
NONMEM provides a utility routine "format_draws.f90" to format the the draw files for future csv parsing. So by building and running the standalone tool on the .ext file
|
|
one obtains a new w15.ext.csv file with the original draw but
title and summary lines commented out:
|
|
Then we can read this file in R:
|
|
Alternatively, one can read the original .ext file and skip the title and summary lines:
|
|
Note that the "-6" argument here is ad hoc, in that the raw output
files may contain a different number of summary and diagnostic lines.
Alternatively one can simply remove the
lines with ITERATION<=-1000000000.
Now we can do postprocessing through the R package posterior. We first
read in the data object as draws. We also skip the warmup (burn-in)
draws, the ones with ITERATION < 0 (the code shows ligatures so it may
not be the same as how it is typed, e.g. the R pipe "|>" is shown as
triangle |>).
|
|
We first examine the summary of the draws.
|
|
Note that the file read maps index notation,
e.g. "SIGMA(1,1)" to "SIGMA.1.1." and "PHI(1)" to "PHI.1.".
The standard deviation in the summary is exactly what is reported on
dummy ITERATION "-1000000001" in the original raw output. Details of
rhat, ess_bulk, and ess_tail can be found here.
We can visulize the draws using the bayesplot package. For example,
|
|
plots the posterior desity,
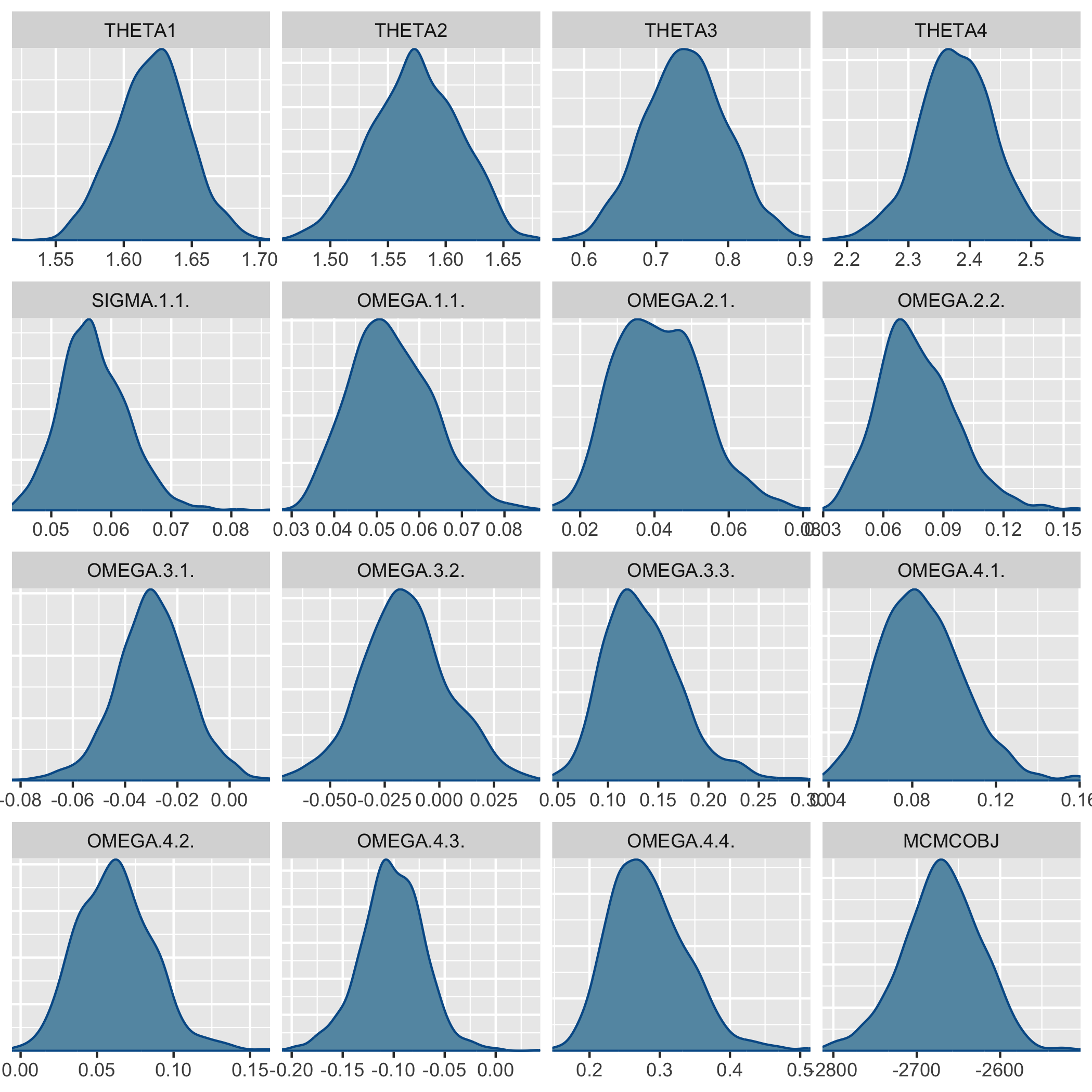
and
|
|
visulizes the correlation between THETAs (regex_pars option selects
the parameters by regular expression, here all the THETA variables). We can see that the
clearance and the volume of distribution V1 & V2 are weakly correlated.
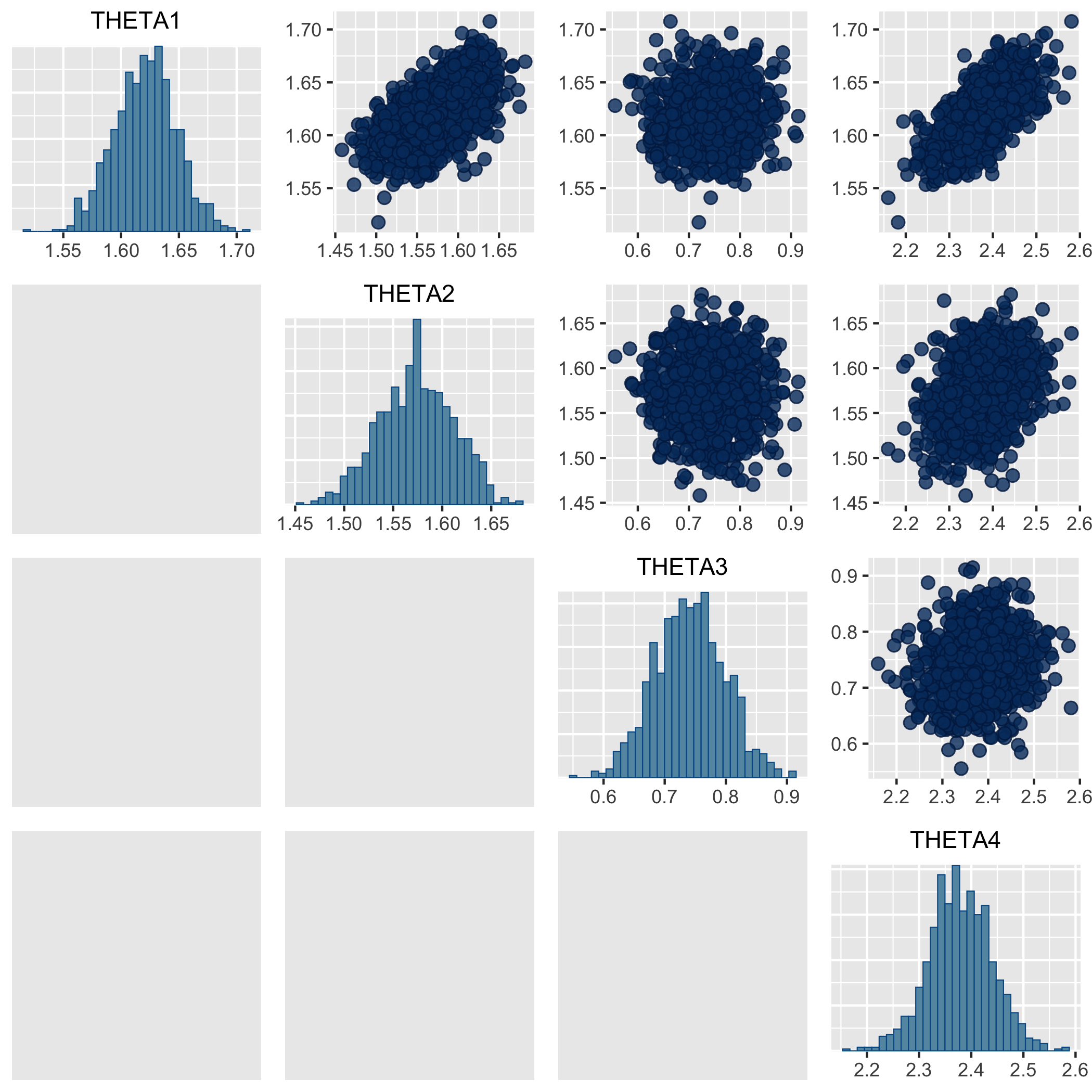
With option BAYES_PHI_STORE=1, the individual draws are also output
in the .iph file. We can follow the same steps to read in those draws.
Note that .iph file does not contain the summary lines.
|
|
This draw object is much bigger than the population parameters, as it contains the draws of PHIs and ETAs for each of the 100 subjects.
The posterior package provides many functions to manipulate draws. For
example, we can extract the individual parameters for subject 1, and
bind_draws with the population parameter draws.
|
|
The following pair plot shows that PHI and ETA are indeeded correlated, and the residual error is likely not associated with them.
|
|
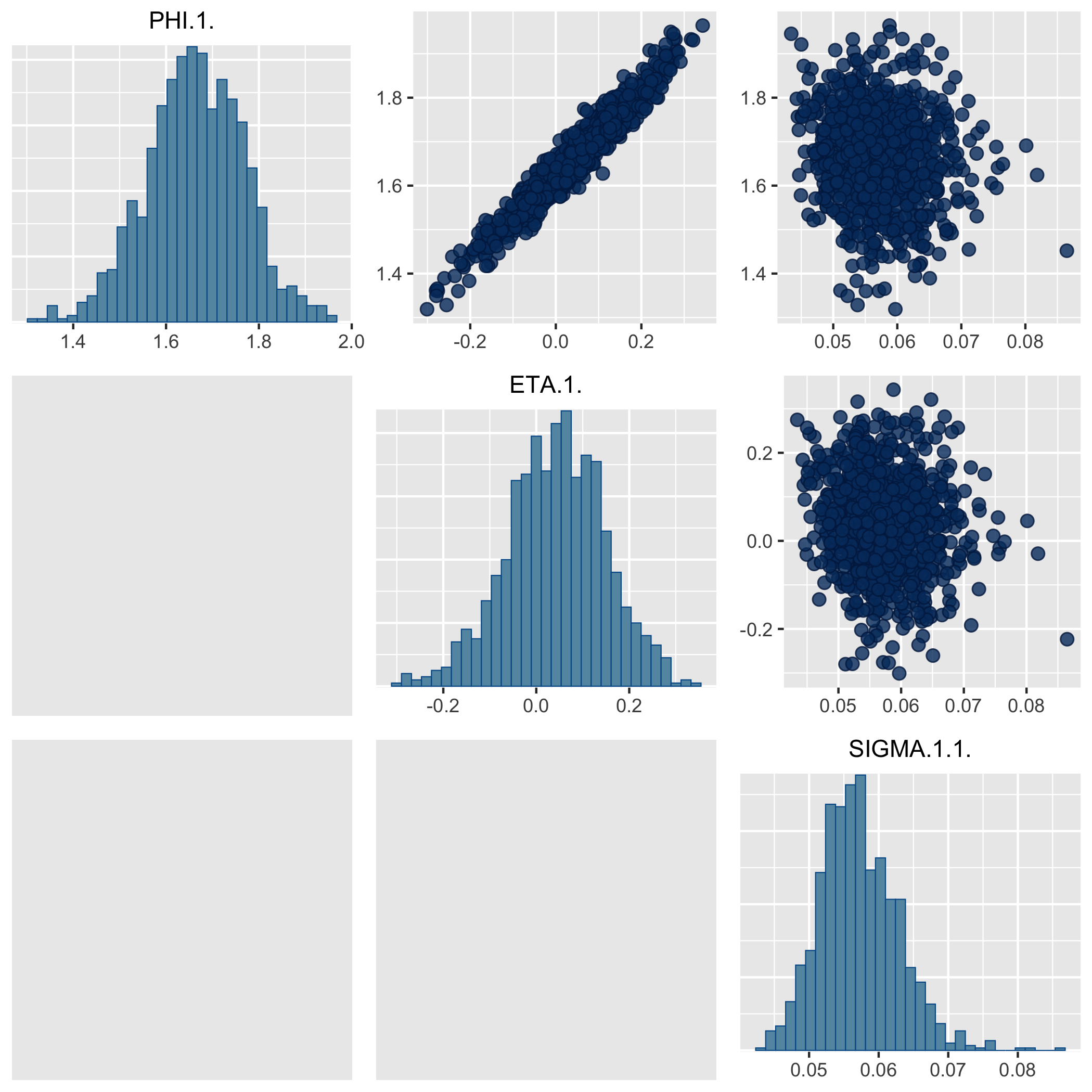
We encourage users to explore the mentioned R packages for additional features.